Search Count: 34
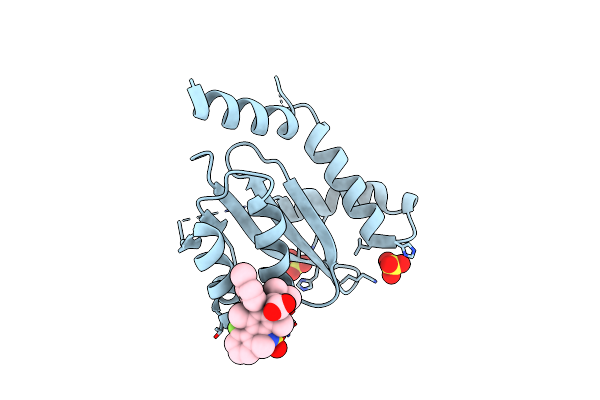 |
Crystal Structure Of Hiv-1 Integrase Catalytic Core Domain In Complex With (2S)-2-(Tert-Butoxy)-2-(10-Fluoro-2-(2-Hydroxy-4-Methylphenyl)-1,4-Dimethyl-5-(Methylsulfonyl)-5,6-Dihydrophenanthridin-3-Yl)Acetic Acid
Organism: Human immunodeficiency virus type 1 (new york-5 isolate)
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2022-09-07 Classification: HYDROLASE Ligands: SO4, 8Z3, GOL |
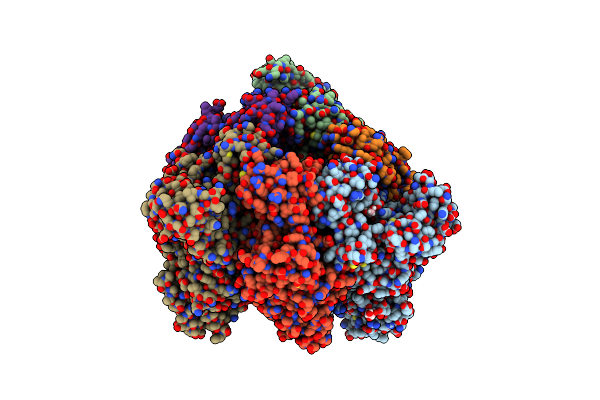 |
Crystal Structure Of Nucleotide-Free Mutant A(S23C)3B(N64C)3 Complex From Enterococcus Hirae V-Atpase
Organism: Enterococcus hirae (strain atcc 9790 / dsm 20160 / jcm 8729 / lmg 6399 / nbrc 3181 / ncimb 6459 / ncdo 1258)
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2021-12-29 Classification: HYDROLASE Ligands: GOL |
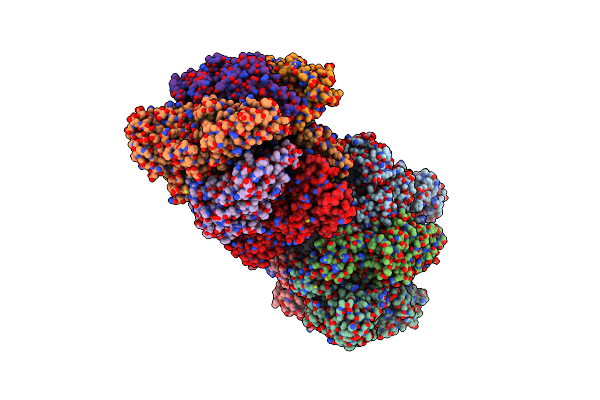 |
Crystal Structure Of The Amp-Pnp-Bound Mutant A(S23C)3B(N64C)3 Complex From Enterococcus Hirae V-Atpase
Organism: Enterococcus hirae (strain atcc 9790 / dsm 20160 / jcm 8729 / lmg 6399 / nbrc 3181 / ncimb 6459 / ncdo 1258)
Method: X-RAY DIFFRACTION Resolution:3.38 Å Release Date: 2021-12-29 Classification: HYDROLASE Ligands: GOL, ANP, MG |
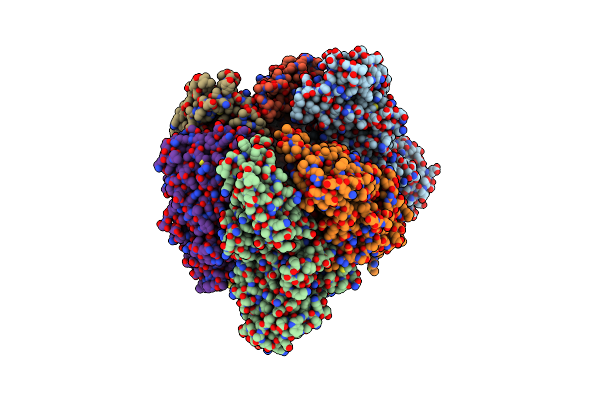 |
Crystal Structure Of The Adp-Bound Mutant A(S23C)3B(N64C)3 Complex From Enterococcus Hirae V-Atpase
Organism: Enterococcus hirae (strain atcc 9790 / dsm 20160 / jcm 8729 / lmg 6399 / nbrc 3181 / ncimb 6459 / ncdo 1258)
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2021-12-29 Classification: HYDROLASE Ligands: MG, ADP, SO4 |
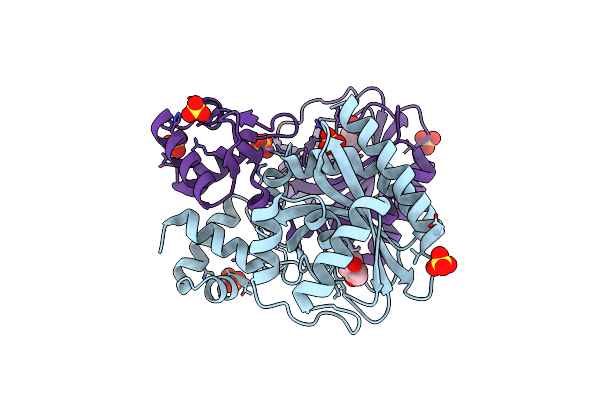 |
Iclr Transcription Factor Complexed With 4-Hydroxybenzoic Acid From Microbacterium Hydrocarbonoxydans
Organism: Microbacterium hydrocarbonoxydans
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-05-05 Classification: GENE REGULATION Ligands: SO4, PHB |
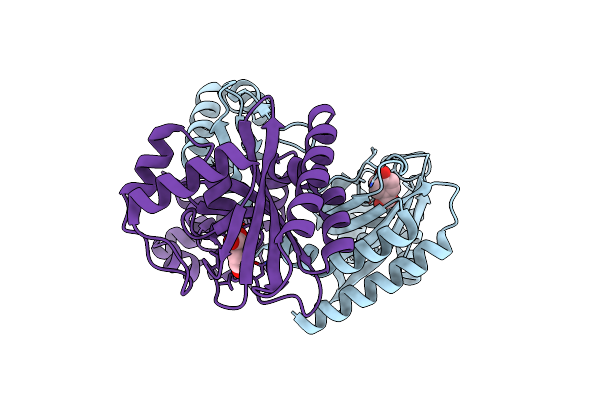 |
Crystal Structure Of An Iclr Homolog Complexed With 4-Hydroxybenzoate From Microbacterium Hydrocarbonoxydans In P212121 Form
Organism: Microbacterium hydrocarbonoxydans
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2021-05-05 Classification: TRANSCRIPTION Ligands: PHB |
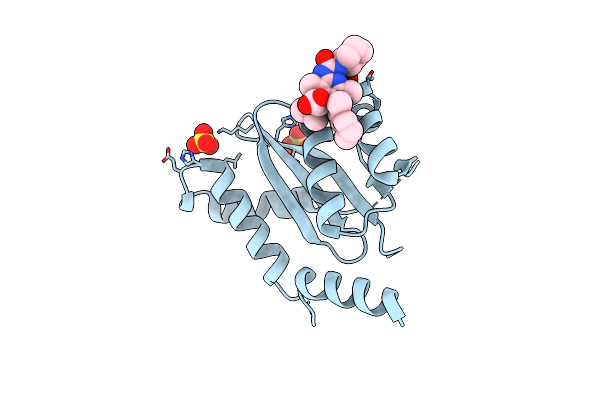 |
Crystal Structure Of Hiv-1 Integrase Catalytic Core Domain In Complex With 2-(Tert-Butoxy)-2-(2-(3-Cyclohexylureido)-3,6-Dimethyl-5-(5-Methylchroman-6-Yl)Pyridin-4-Yl)Acetic Acid
Organism: Human immunodeficiency virus type 1 group m subtype b (isolate ny5)
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: SO4, GZ9 |
 |
Organism: Rubrobacter xylanophilus (strain dsm 9941 / nbrc 16129)
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2020-04-08 Classification: TRANSPORT PROTEIN Ligands: MPG, RET, SO4 |
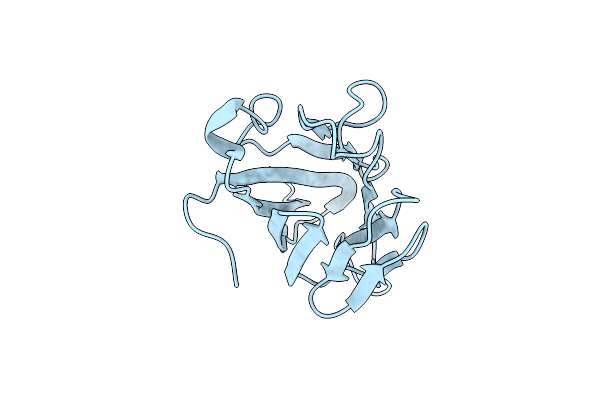 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2020-04-01 Classification: SIGNALING PROTEIN |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2020-04-01 Classification: SIGNALING PROTEIN |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2020-04-01 Classification: SIGNALING PROTEIN |
 |
Crystal Structure Of A Substrate Binding Protein From Microbacterium Hydrocarbonoxydans
Organism: Microbacterium hydrocarbonoxydans
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2020-03-25 Classification: TRANSPORT PROTEIN |
 |
Crystal Structure Of A Substrate Binding Protein From Microbacterium Hydrocarbonoxydans Complexed With 4-Hydroxybenzoate Hydrazide
Organism: Microbacterium hydrocarbonoxydans
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2020-03-25 Classification: TRANSPORT PROTEIN Ligands: HDH |
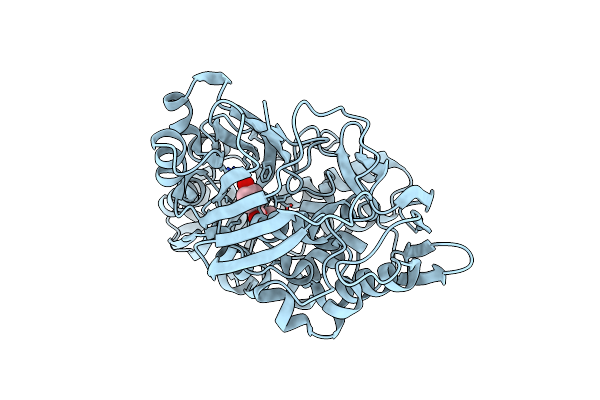 |
Crystal Structure Of The Substrate Binding Protein From Microbacterium Hydrocarbonoxydans Complexed With Propylparaben
Organism: Microbacterium hydrocarbonoxydans
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-03-25 Classification: TRANSPORT PROTEIN Ligands: 36M |
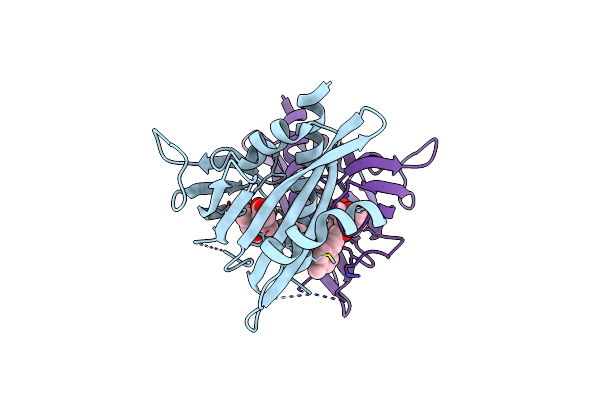 |
Crystal Structure Of The Abscisic Acid Receptor Pyr1 In Complex With An Antagonist
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2018-04-18 Classification: HORMONE RECEPTOR Ligands: 8V6 |
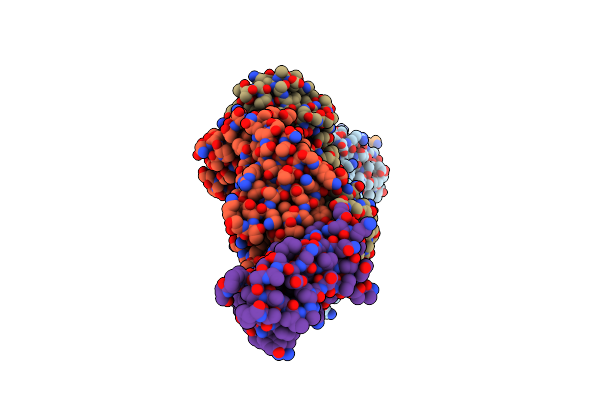 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2017-05-03 Classification: IMMUNE SYSTEM |
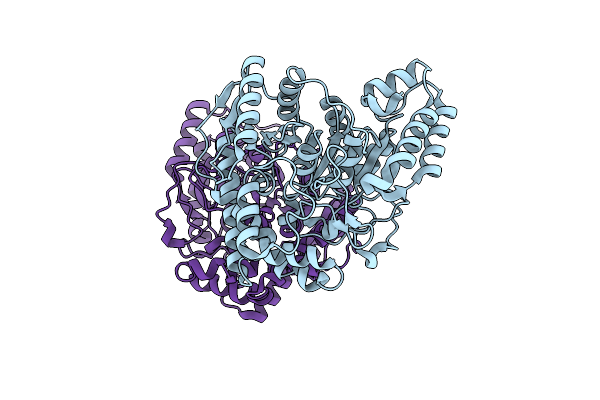 |
Organism: Microbacterium sp. hm58-2
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2017-03-15 Classification: HYDROLASE |
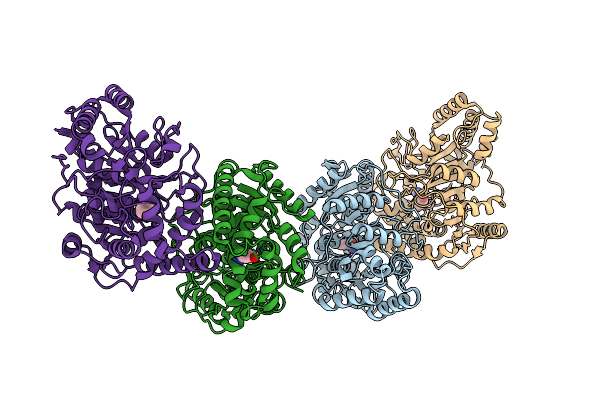 |
Organism: Microbacterium sp. hm58-2
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2017-02-01 Classification: HYDROLASE Ligands: HDH |
 |
Organism: Microbacterium sp.
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2017-01-18 Classification: TRANSCRIPTION REGULATOR Ligands: PO4 |
 |
Crystal Structure Of Ha33 From Clostridium Botulinum Serotype C Strain Yoichi
Organism: Clostridium botulinum
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2016-06-15 Classification: SUGAR BINDING PROTEIN Ligands: PGE |

