Search Count: 73
 |
Crystal Structure Of The Glycosyltransferase Domain Of Legionella Seta (Unbound)
Organism: Legionella pneumophila
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2025-02-19 Classification: TRANSFERASE Ligands: MG |
 |
Crystal Structure Of The Glycosyltransferase Domain Of Legionella Seta In Complex With Updglc
Organism: Legionella pneumophila
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-02-19 Classification: TRANSFERASE Ligands: UDP, MG, BGC, SO4 |
 |
Crystal Structure Of The Glycosyltransferase Domain Of Legionella Seta In Complex With Upd
Organism: Legionella pneumophila
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2025-02-19 Classification: TRANSFERASE Ligands: UDP, MG, EDO, GOL |
 |
Crystal Structure Of The Pi3P-Binding Domain Of Legionella Seta In Complex With Inositol 1,3-Bisphosphate
Organism: Legionella pneumophila
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2025-02-19 Classification: TRANSFERASE Ligands: GOL, ITP |
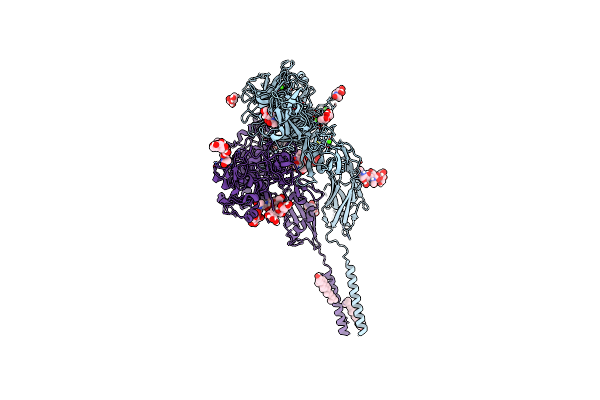 |
Cryo-Em Structures Of Full-Length Integrin Alphaiibbeta3 In Native Lipids Complexed With Modified Tirofiban
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION Ligands: CA, NAG, MG, A1A5G, CLR |
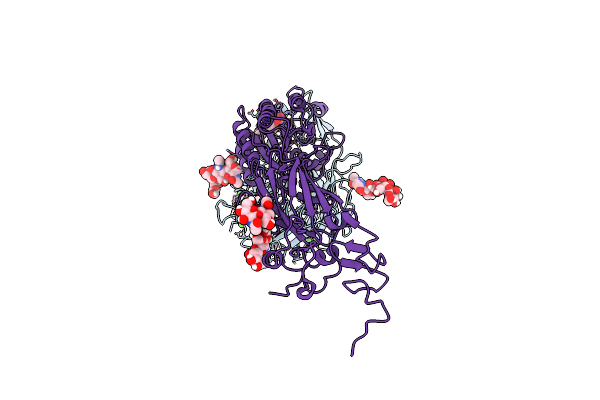 |
Cryo-Em Structures Of Full-Length Integrin Alphaiibbeta3 In Native Lipids Complexed With Tirofiban
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION Ligands: CA, MG, AGG |
 |
Crystal Structure Of Human Methionine Aminopeptidase 12 (Map12) In The Unbound Form
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2024-01-24 Classification: HYDROLASE Ligands: PGE, GOL |
 |
Crystal Structure Of Human Methionine Aminopeptidase 12 (Map12) In Complex With Two Cobalt Ions
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2024-01-24 Classification: HYDROLASE Ligands: CO, PGE |
 |
Crystal Structure Of Human Methionine Aminopeptidase 12 (Map12) In Complex With Two Cobalt Ions And Methionine
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-01-24 Classification: HYDROLASE Ligands: MET, CO, PGE |
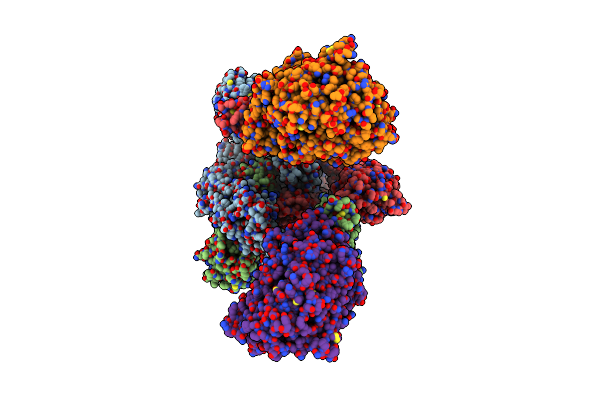 |
Cryo-Em Structure Of Mink Variant Y453F Trimeric Spike Protein Bound To Two Mink Ace2 Receptors
Organism: Severe acute respiratory syndrome coronavirus 2, Neovison vison
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: VIRAL PROTEIN |
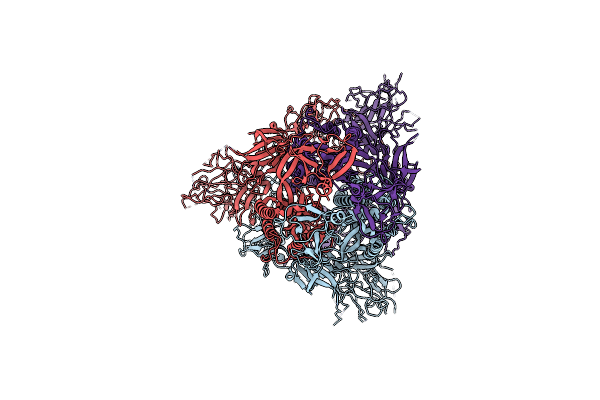 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: VIRAL PROTEIN |
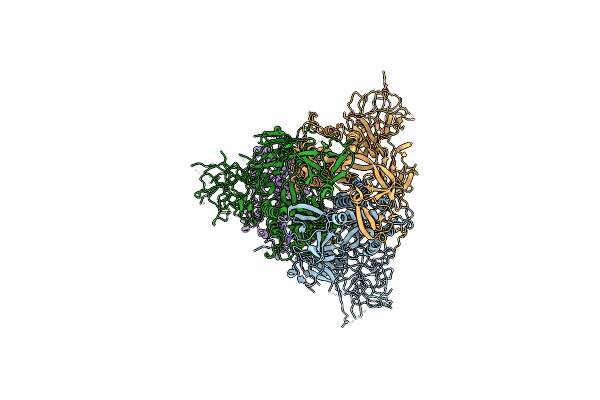 |
Cryo-Em Structure Of Mink Variant Y453F Trimeric Spike Protein Bound To One Mink Ace2 Receptors At Downrbd Conformation
Organism: Severe acute respiratory syndrome coronavirus 2, Neovison vison
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: VIRAL PROTEIN |
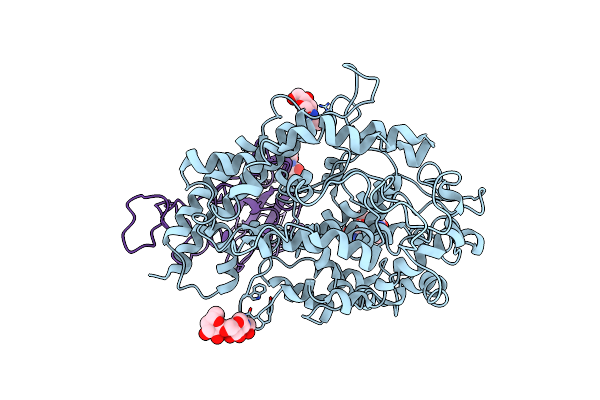 |
Cryo-Em Structure Of The Rbd-Ace2 Interface Of The Sars-Cov-2 Trimeric Spike Protein Bound To Ace2 Receptor After Local Refinement At Uprbd Conformation
Organism: Neovison vison, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: VIRAL PROTEIN Ligands: NAG |
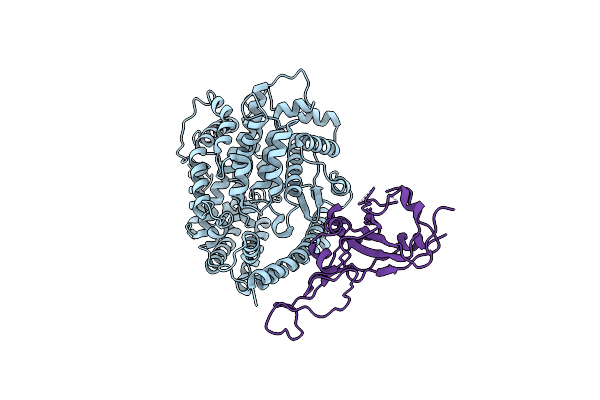 |
Cryo-Em Structure Of The Rbd-Ace2 Interface Of The Sars-Cov-2 Trimeric Spike Protein Bound To Ace2 Receptor After Local Refinement At Downrbd Conformation.
Organism: Neovison vison, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: VIRAL PROTEIN |
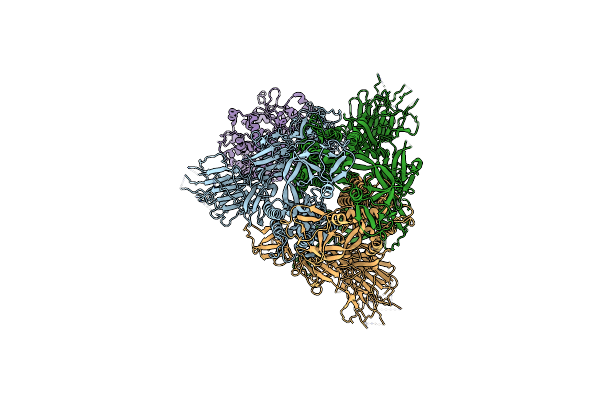 |
Cryo-Em Structure Of Mink Variant Y453F Trimeric Spike Protein Bound To One Mink Ace2 Receptors
Organism: Severe acute respiratory syndrome coronavirus 2, Neogale vison
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: VIRAL PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.48 Å Release Date: 2023-07-26 Classification: TRANSFERASE Ligands: J6F |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-11-09 Classification: LYASE Ligands: PO4, MG, ACT, EDO, PGE, SEP |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2022-02-23 Classification: METAL BINDING PROTEIN Ligands: TRS, CL, NI, ZN |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2022-02-23 Classification: METAL BINDING PROTEIN Ligands: CMP, TRS, CL, ZN, NI |
 |
Crystal Structure Of Recombinant Human Fumarase In Complex With D-2-Amino-3-Phosphono-Propionic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2022-02-23 Classification: LYASE Ligands: APO, GOL |

