Search Count: 917
 |
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: EDO, ZN |
 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn From Cupriavidus Pinatubonensis In Complex With Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI |
 |
Crystal Structure Of Hpsn D352A Mutant From Cupriavidus Pinatubonensis In Complex With Nad+
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: NAD, ZN |
 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn From Cupriavidus Pinatubonensis In Complex With Product Analogue (L-Cysteate) And Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI, OCS |
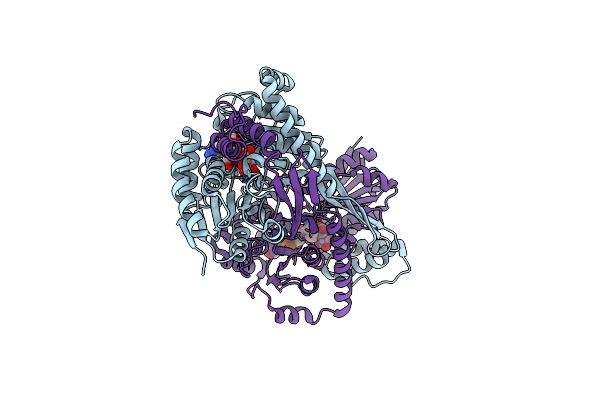 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn From Cupriavidus Pinatubonensis In Complex With Product (R-Sulfolactate) And Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI, 3SL |
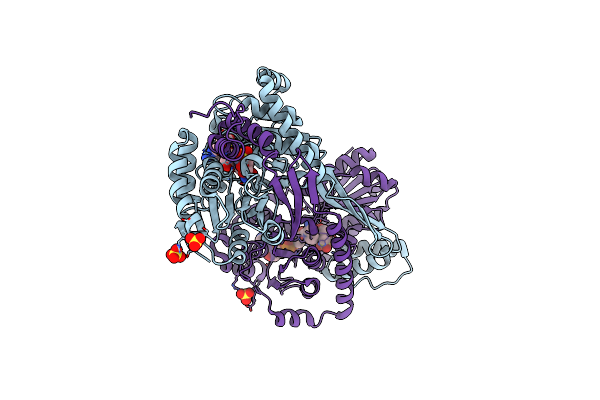 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn H319A Variant From Cupriavidus Pinatubonensis In Complex With Substrate (R-Dhps) And Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI, A1AZH, SO4 |
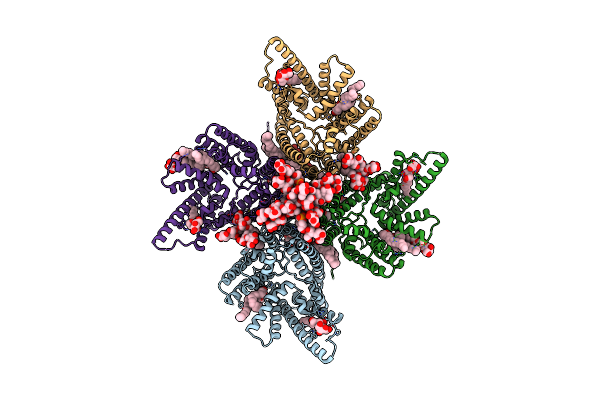 |
Organism: Streptococcus thermophilus lmg 18311
Method: ELECTRON MICROSCOPY Release Date: 2024-05-22 Classification: MEMBRANE PROTEIN Ligands: PGT, LMT, DGA |
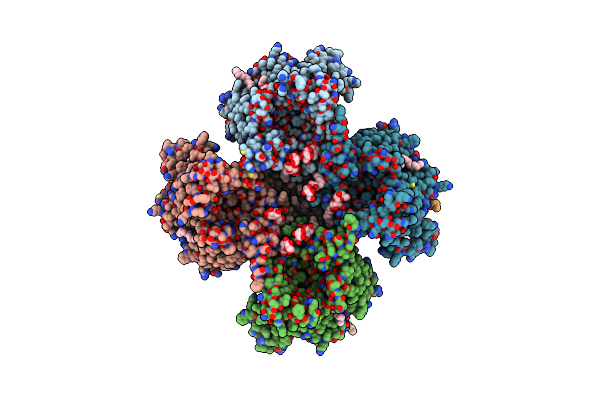 |
Organism: Streptococcus thermophilus lmg 18311
Method: ELECTRON MICROSCOPY Release Date: 2024-05-22 Classification: MEMBRANE PROTEIN Ligands: PGT, LMT, DGA |
 |
Organism: Streptococcus thermophilus lmg 18311
Method: ELECTRON MICROSCOPY Release Date: 2024-05-15 Classification: MEMBRANE PROTEIN/INHIBITOR Ligands: ASW, PGT, LMT, DGA |
 |
Structure Of Sponge-Phase Grown Peptst2 Collected By Rotation Serial Crystallography On A Coc Membrane At A Synchrotron Source
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311)
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2021-09-01 Classification: TRANSPORT PROTEIN Ligands: PO4, 78M, PG0, PGE |
 |
Organism: Streptococcus thermophilus lmg 18311
Method: X-RAY DIFFRACTION Resolution:2.96 Å Release Date: 2021-07-21 Classification: TRANSFERASE Ligands: SO4 |
 |
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311)
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2021-04-21 Classification: HYDROLASE Ligands: GDP, ALF, MG, K, CL, PEG, EDO |
 |
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2021-04-14 Classification: HYDROLASE Ligands: EDO, GDP, ALF, MG, K, CL |
 |
Structure Of Peptst From Coc Imisx Setup Collected By Rotation Serial Crystallography On Crystals Prelocated By 2D X-Ray Phase-Contrast Imaging
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311)
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2020-11-04 Classification: TRANSPORT PROTEIN Ligands: PO4, 78M, PG0, PGE |
 |
Structure Of Peptst From Coc Imisx Setup Collected By Still Serial Crystallography On Crystals Prelocated By 2D X-Ray Phase-Contrast Imaging
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311)
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2020-11-04 Classification: TRANSPORT PROTEIN Ligands: PO4, 78M, PG0, PGE |
 |
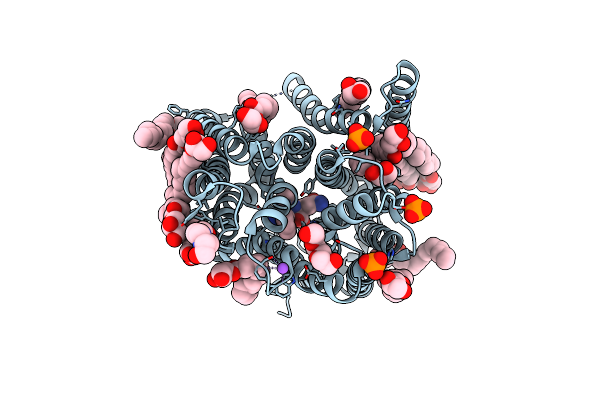
Crystal Structure Of Streptococcus Thermophilus Shp Pheromone Receptor Rgg3
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311)
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2020-10-21 Classification: DNA BINDING PROTEIN Ligands: SO4 |
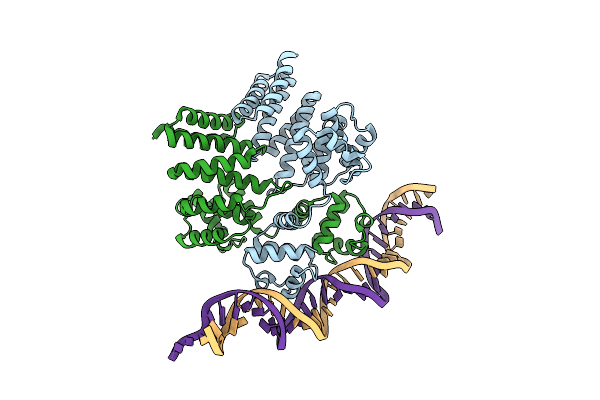 |
Crystal Structure Of Streptococcus Thermophilus Shp Pheromone Receptor Rgg3 Bound To Dna
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311), Streptococcus thermophilus cnrz1066
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2020-10-21 Classification: DNA BINDING PROTEIN/DNA |
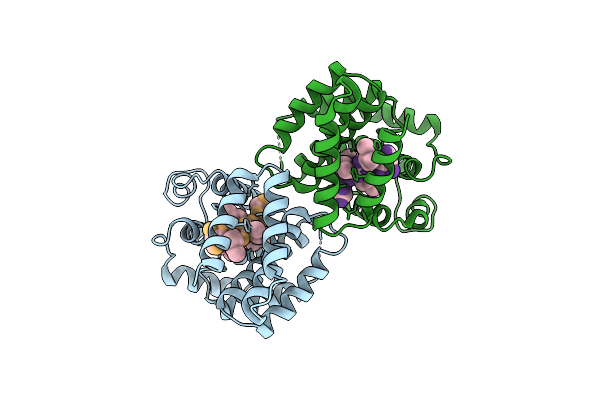 |
Cryoem Structure Of Streptococcus Thermophilus Shp Pheromone Receptor Rgg3 In Complex With Shp3
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311), Streptococcus thermophilus cnrz1066
Method: ELECTRON MICROSCOPY Release Date: 2020-10-07 Classification: DNA BINDING PROTEIN |
 |
Organism: Streptococcus thermophilus (strain atcc baa-250 / lmg 18311), Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2018-09-19 Classification: MEMBRANE PROTEIN Ligands: PO4, 1PE, EPE, NA, 78N, 78M |
 |
Organism: Escherichia coli m718, Escherichia coli m605, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.70 Å Release Date: 2018-09-05 Classification: TRANSCRIPTION/RNA Ligands: ADP, MG, BEF |

